The Past and The Future
The Past
I recently finished my Master thesis and can now call myself Master of Science. The thesis itself couldn’t turn out any better as it did. The goal was to implement a method that is capable to predict the relative location along the longitudinal axis of a slice of a X-ray computed tomography (CT) volume. The motivation behind this is to speed up the process when a physician wants to compare two CT volumes against each other. Usually, the patient’s data can be accessed over the network and a single CT volume can contain more than thousand images taking more than 1 GB disk space. In addition, most physicians are only interested in one particular area they want to compare. Accordingly, it would be waste of resources if one has to transfer and load two complete volumes and manually navigate to the area of interest. The methods developed in this thesis allow the physician to select the area of interest (on the longitudinal axis) in one volume and only transfer and load the corresponding area in the other volume. Obviously, you get the biggest benefit if the volume contains a large amount of images and the region of interest is relatively small.
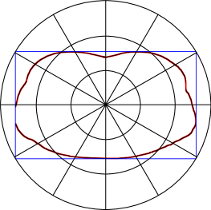
Histogram in polar space.
Pixel values in CT images are reliable and are measured in Hounsfield units, where 0 denotes water, highly negative values air, and values greater than 300 bone structures. Based on these intensities it is easy to find all pixels that represent air or bones. For each image two different histograms in polar space where created. The first one counts the pixels representing bone structures and the second pixels that represent air inclusions inside the body. In the end a single image was be described by 344 features.
Prediction is based on instance-based regression using k-nearest neighbor search. Therefore, It depends on a database of CT volumes with annotated positions for each image. For a query image the feature vector described above is extracted and used to search the database for the three nearest neighbors to this vector using the cosine distance. The predicted position is calculated by averaging the annotated positions of all nearest neighbors. In the end tests showed that the mean prediction error is approximately 16 mm, which is a 2.5-fold improvement compared to state of the art methods on the same dataset.
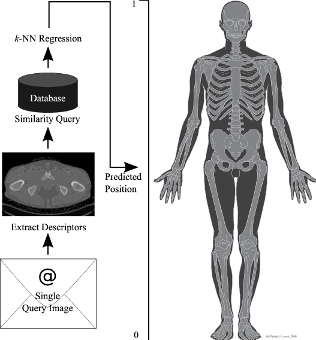
Instance-based regression scheme.
Finally, the results were so convincing that my work was the basis for two publications, which was the icing on the cake. The first one named Position Prediction in CT Volume Scans was presented at the ICML 2011 Workshop on Machine Learning for Global Challenges, and the second one named 2D Image Registration in CT Images using Radial Image Descriptors will be presented at this years MICCAI conference. Accordingly, I couldn’t be happier about my thesis.
The Future
As much as I enjoyed the last five years studying in Munich the “real” world is waiting. Therefore, I’m currently looking for a position as software developer. I spent over a year working for my university’s Database Systems Group, which allowed me to build extensive experience and interest in machine learning, data mining and databases. The ideal position would be related to these areas. In addition, I have an increasing interest in dealing with large data, i.e. extremely large databases and/or high dimensional data and massively parallel computing. My current work at the database systems group involves coming up with improved strategies (pruning, on-disk structure) to speed up k-nearest neighbor search for high dimensional data similar to an earlier publication.
If you are working in some of the areas mentioned above or know of interesting job openings, feel free to contact me. You can have a look at my CV, as well.